| Content: | 100 / 200 assays per kit |
| Shipping Temperature: | Ambient |
| Storage Temperature: |
Short term stability: 2-8oC, Long term stability: See individual component labels |
| Stability: | > 2 years under recommended storage conditions |
| Analyte: | Pullulanase/Limit-Dextrinase |
| Assay Format: | Spectrophotometer |
| Detection Method: | Absorbance |
| Wavelength (nm): | 400 |
| Signal Response: | Increase |
| Limit of Detection: |
0.18 U/mL for pullulanase preparations (50-fold dilution) 0.01 U/g for limit dextrinase in milled malt |
| Reproducibility (%): | ~ 3% |
| Total Assay Time: |
~ 10 min (Pullanase), ~ 30 min (Limit-Dextrinase) |
| Application examples: | Assay of microbial pullulanase preparations. Measurement of limit-dextrinase in malt extracts. |
| Method recognition: | Novel method |
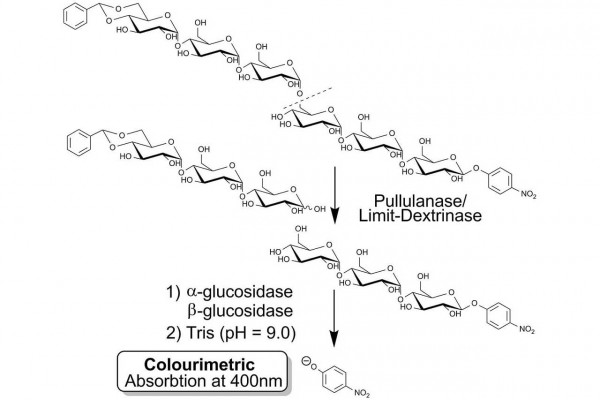
PullG6 assay for the measurement of pullulanase employs a water soluble defined substrate, namely 4,6-O-benzylidene-4-nitrophenyl-63-α-D-maltotriosyl-maltotriose (BPNPG3G3), coupled with the ancillary enzymes α-glucosidase and β-glucosidase. Upon hydrolysis of the substrate at the 1,6-α-linkage by pullulanase or limit-dextrinase, the released 4-nitrophenyl-β-maltotrioside is immediately hydrolysed to glucose and 4-nitrophenol by the concerted action of the α-glucosidase and β-glucosidase enzymes in the reagent mixture. The reaction is terminated and phenolate ions are developed by addition of dilute alkali. The absorbance is read at 400 nm and the value obtained correlates directly with pullulanase activity.
Explore more of our assay kit products for enzyme activity measurement.

- High sensitivity
- Suitable for manual and auto-analyser formats
- No transglycosylation interference
- Very cost effective
- All reagents stable for > 1 year after preparation
- Very specific
- Simple format
- Standard included
Diastatic power and maltose value: a method for the measurement of amylolytic enzymes in malt.
Charmier, L. M., McLoughlin, C. & McCleary, B. V. (2021). Journal of the Institute of Brewing, In Press.
A simple method for measurement of the amylolytic activity of malt has been developed and fully evaluated. The method, termed the Maltose Value (MV) is an extension of previously reported work. Here, the MV method has been studied in detail and all aspects of the assay (sample grinding and extraction, starch hydrolysis, maltose hydrolysis and determination as glucose) have been optimised. The method is highly correlated with other dextrinising power methods. The MV method involves extraction of malt in 0.5% sodium chloride at 30°C for 20 minutes followed by filtration; incubation of an aliquot of the undiluted filtrate with starch solution (pH 4.6) at 30°C for 15 min; termination of reaction with sodium hydroxide solution; dilution of sample in an appropriate buffer; hydrolysis of maltose with a specific α-glucosidase; glucose determination and activity calculation. Unlike all subsequent reducing sugar methods, the maltose value method measures a defined reaction product, maltose, with no requirement to use equations to relate analytical values back to Lintner units. The maltose value method is the first viable method in 130 years that could effectively replace the 1886 Lintner method.
Hide AbstractPrediction of potential malt extract and beer filterability using conventional and novel malt assays.
Cornaggia, C., Evans, D. E., Draga, A., Mangan, D. & McCleary, B. V. (2019). Journal of Institute of Brewing, 125(3), 294-309.
Colourimetric assays were used to measure the activities of six key hydrolases endogenous to barley: β‐glucanase, xylanase, cellulase, α-amylase, beta‐amylase and limit dextrinase. The analysed barley malt samples were previously characterised by 27 conventional malt quality descriptors. Correlations between enzymatic activities and brewing parameters such as extract yield, fermentability, viscosity and filterability were investigated. A single extraction protocol for all six hydrolases was optimised and used for multi‐enzyme analysis using fully automatable assay formats. A regression analysis between malt parameters was undertaken to produce a relationship matrix linking enzyme activities and conventional malt quality descriptors. This regression analysis was used to inform a multi‐linear regression approach to create predictive models for extract yield, apparent attenuation limit, viscosity and filterability using the Small‐scale Wort rapId Filtration Test (SWIFT) and two different mashing protocols – Congress and a modified infusion mash at 65oC (MIM 65oC). It was observed that malt enzyme activities displayed significant correlations with the analysed brewing parameters. Both starch hydrolases and cell wall hydrolase activities together with modification parameters (i.e. Kolbach index) were found to be highly correlated with extract yield and apparent attenuation limit. Interestingly, it was observed that xylanase activity in malts was an important predictor for wort viscosity and filterability. It is envisaged that the automatable measurement of enzyme activity could find use in plant breeding progeny selection and for routine assessment of the functional brewing performance of malt batches. This analytical approach would also contribute to brewing process consistency, product quality and reduced processing times.
Hide AbstractMangan, D., McCleary, B. V., Cornaggia, C., Ivory, R., Rooney, E. & McKie, V. (2015). Journal of Cereal Science, 62, 50-57.
The measurement of limit-dextrinase (LD) (EC 3.2.1.142) in grain samples such as barley, wheat or rice can be problematic for a number of reasons. The intrinsic LD activity in these samples is extremely low and they often contain a limit-dextrinase inhibitor and/or high levels of reducing sugars. LD also exhibits transglycosylation activity that can complicate the measurement of its hydrolytic activity. A minor modification to the industrial standard Limit-Dextrizyme tablet test is suggested here to overcome this transglycosylation issue.
Hide AbstractMcCleary, B. V., Mangan, D., McKie, V., Cornaggia, C., Ivory, R. & Rooney, E. (2014). Carbohydrate Research, 393, 60-69.
Specific and highly sensitive colourimetric and fluorometric substrate mixtures have been prepared for the measurement of pullulanase and limit-dextrinase activity and assays employing these substrates have been developed. These mixtures comprise thermostable α- and β-glucosidases and either 4,6-O-benzylidene-2-chloro-4-nitrophenyl-β-maltotriosyl (1-6) α-maltotrioside (BzCNPG3G3, 1) as a colourimetric substrate or 4,6-O-benzylidene-4-methylumbelliferyl-β-maltotriosyl (1-6) α-maltotrioside (BzMUG3G3, 2) as a fluorometric substrate. Hydrolysis of substrates 1 and 2 by exo-acting enzymes such as amyloglucosidase, β-amylase and α-glucosidase is prevented by the presence of the 4,6-O-benzylidene group on the non-reducing end D-glucosyl residue. The substrates are not hydrolysed by any α-amylases studied, (including those from Aspergillus niger and porcine pancreas) and are resistant to hydrolysis by Pseudomonas sp. isoamylase. On hydrolysis by pullulanase, the 2-chloro-4-nitrophenyl-β-maltotrioside (3) or 4-methylumbelliferyl-β-maltotrioside (4) liberated is immediately hydrolysed to D-glucose and 2-chloro-4-nitrophenol or 4-methylumbelliferone. The reaction is terminated by the addition of a weak alkaline solution leading to the formation of phenolate ions in solution whose concentration can be determined using either spectrophotometric or fluorometric analysis. The assay procedure is simple to use, specific, accurate, robust and readily adapted to automation.
Hide AbstractDiastatic Activity of German Hop Cultivars with Respect to Variety, Crop Year, and Separated Hop Cone Parts.
Wietstock, P. C., Michalek, D., Treetzen, T., Pinto, M. B. C., Biendl, M. & Gibson, B. (2025). ACS Food Science & Technology, 5(6), 2408-2416.
Dry hopping of beer can result in unintended refermentation, also known as hop creep, because of intrinsic hop diastatic activity. The objective of the work described herein was to determine the enzymatic activity across 16 different hop cultivars grown in Germany in crop years 2019, 2020, and 2021. Optimized enzyme kit protocols were used to quantitate hop α-, β-amylase, amyloglucosidase, and limit dextrinase activities, while a recently published method measured hop diastatic activity. Clear varietal distinctions exist, and hops of harvests 2019, 2020, and 2021 were subsequently classified into three groups depending on their enzymatic activity. With respect to different harvest years, the results imply an annual influence on the amylolytic activity of hops in principle, but more monitoring is needed. Processing methods such as pelletization and storage under different conditions showed a minimal impact on enzymatic activity. Based on further sampling from hops of the harvest 2022, it was observed that differences among hop fractions are pronounced, with the vegetative material and strig exhibiting higher enzymatic activity compared to the lupulin fraction.
Hide AbstractA quick and simple gel diffusion assay to visualize and quantify pullulanase activity for resistant starch content in food crops.
Krishnan, V., Awana, M., Kulshreshta, A., Praveen, S. & Singh, A. (2022). Journal of Plant Biochemistry and Biotechnology, 1-7.
Pullulanases (PULs) are glucanohydrolases which break α-1,6 glycosidic bonds. They have immense potential in food industry to develop low calorie foods and hence new biological sources needs to be identified. Till date PUL enzyme has been assayed with fluorogenic/radiolabeled substrates, while the present study aimed to develop a non-radioactive, quick, simple, and inexpensive gel-diffusion assay for PUL screening. Here, PUL hydrolyzes chromogenic substrate (red pullulan) leaving a hydrolytic zone alongwith change in intensity based on its activity. Response variables like substrate concentrations (0.5-2% red pullulan), enzyme concentrations (0.0125-0.4 U) and gel depth (0.25-1.5 cm) were optimized for maximum efficiency. Seeded agarose gel depth of 1 cm with 1% red pullulan detected lowest limit of detection (LOD) of PUL activity (0.0125 U). Based on the positive correlation between PUL activity and resistant starch (RS) content, the developed tool was validated using contrasting RS rice genotypes, which widened the scope of this tool for screening high RS cultivars in future. This will give a way to breeders for developing nutritionally superior, high RS content rice varieties.
Hide AbstractEffects of post-heading high temperature on some quality traits of malt barley.
Ni, S. J., Zhao, H. F. & Zhang, G. P. (2020). Journal of Integrative Agriculture, 19(11), 2674-2679.
Global change is bringing barley with more frequency of suffering from high temperature. However, little has been known about the influence of high temperature on malt quality traits. In this study, we investigated the impact of 1-wk heat stress (32°C/26°C, day/night, 12 h/12 h) initiating from the 7th (HT7) and 14th (HT14) days after heading on some grain and malt quality traits of two barley cultivars. In comparison with normal temperature (24°C/18°C, day/night, 12 h/12 h), heat stress significantly reduced kernel weight, seed setting rate and grains per spike: HT7 having a larger effect than HT14. Meanwhile, total protein and β-glucan contents, and β-amylase and limit dextrinase activities were significantly increased under high temperature, with HT7-treated plants showing larger changes. Moreover, the different changes of four protein fractions under heat stress were found in the two barley cultivars, indicating the possibility of breaking positive association between protein content and enzyme activity.
Hide AbstractHigh amylose wheat starch structures display unique fermentability characteristics, microbial community shifts and enzyme degradation profiles.
Bui, A., Williams, B., Hoedt, E., Morrison, M., Mikkelsen, D. & Gidley, M. (2020). Food & Function, 6.
A slower rate of starch digestion in the small intestine increases the amount of resistant starch (RS) entering the large intestine, which is associated with health benefits. Although increasing the amylose (AM) content of dietary starch intake is one way to increase RS, the processes involved in gut microbial hydrolysis and fermentation of high AM-RS substrates are poorly understood. In this study, five high AM wheat (HAW) starches ranging from 47% AM to 93% AM and a wild type (37% AM), in both native granular and cooked forms, were subjected to in vitro fermentation with a porcine faecal inoculum. Fermentation kinetics, temporal microbial changes, amylolytic enzyme activities and residual starch were determined. All granular starches showed similar fermentation characteristics, independent of AM level, whereas cooking accelerated fermentation of lower AM but slowed fermentation of high AM starches. HAW starches with a very high AM content (>85%) all had similar fermentation kinetics and short-chain fatty acid end-product profiles. Microbial α-amylase, β-amylase, pullulanase and amyloglucosidase enzymatic activities were all detected and followed fermentation kinetics. HAW starch promoted shifts in the microbial community, with increases of the family Lachnospiraceae and the genus Treponema observed, while the genera Prevotella and Streptococcus were reduced in comparison to 37% AM. Overall, these findings suggest that any HAW starch incorporated into high RS food products would be expected to have beneficial microbiota-mediated effects in terms of fermentation kinetics and end products.
Hide AbstractChanges in malt quality during production in two commercial malt houses.
Yousif, A. M. & Evans, D. E. (2020). Journal of the Institute of Brewing, 126 (3), 233-252.
This investigation presents a holistic and comprehensive assessment of the stepwise changes in barley quality during the malting process for multiple batches of two Australian malting varieties (Buloke and Gairdner), in two modern, commercial scale pneumatic malthouses. The study sought to analyse and compare malting plant and variety with respect to basic changes in malt quality for protein (total protein and free amino nitrogen), fermentability (apparent attenuation limit and diastatic power), extract yield, along with filtration indicators (lautering efficiency, viscosity and β‐glucan). Overall, comparing the two malt plants, it was observed that although malt batches and varieties followed different malting pathways, the finished and kilned malt was of satisfactory quality in terms of FAN, viscosity, friability, fermentability and extract.
Hide AbstractThe influence of drought stress on malt quality traits of the wild and cultivated barleys.
Hong, Y. & Zhang, G. P. (2020). Journal of Integrative Agriculture, 19(8), 2009-2015.
As a major abiotic stress, drought causes instability and deterioration of malt barley quality. There is distinct difference among barley cultivars in the responses of the main malt quality traits to drought stress. In the previous study, we identified some Tibetan wild barley accessions with relatively less change of malt quality traits under drought. In this study, we examined the impact of drought stress during grain filling stage on grain weight and several important malt quality traits, including total protein content, β-glucan content, limit dextrinase activity, β-amylase activity, and protein fractions in four barley genotypes (two Tibetan wild accessions and two cultivars). Drought treatment reduced grain weight, β-glucan content, and increased total protein content, β-amylase activity. These changes differed among barley genotypes and treatments, and are closely associated with grain filling process and kernel weight. All the results indicated Tibetan wild barley had great potential for developing drought tolerant barley cultivars. Relatively stable kernel weight or filling process under water stress should be highlighted in malt barley breeding in order to reduce the effect of water stress on malt barley quality.
Hide AbstractNTRC and Thioredoxin f Overexpression Differentially Induces Starch Accumulation in Tobacco Leaves.
Ancín, M., Larraya, L., Millán, F. S., Veramendi, J., Burch-Smith, T. & Farran, I. (2019). Plants, 8(12), 543.
Thioredoxin (Trx) f and NADPH-dependent Trx reductase C (NTRC) have both been proposed as major redox regulators of starch metabolism in chloroplasts. However, little is known regarding the specific role of each protein in this complex mechanism. To shed light on this point, tobacco plants that were genetically engineered to overexpress the NTRC protein from the chloroplast genome were obtained and compared to previously generated Trx f-overexpressing transplastomic plants. Likewise, we investigated the impact of NTRC and Trx f deficiency on starch metabolism by generating Nicotiana benthamiana plants that were silenced for each gene. Our results demonstrated that NTRC overexpression induced enhanced starch accumulation in tobacco leaves, as occurred with Trx f. However, only Trx f silencing leads to a significant decrease in the leaf starch content. Quantitative analysis of enzyme activities related to starch synthesis and degradation were determined in all of the genotypes. Zymographic analyses were additionally performed to compare the amylolytic enzyme profiles of both transplastomic tobacco plants. Our findings indicated that NTRC overexpression promotes the accumulation of transitory leaf starch as a consequence of a diminished starch turnover during the dark period, which seems to be related to a significant reductive activation of ADP-glucose pyrophosphorylase and/or a deactivation of a putative debranching enzyme. On the other hand, increased starch content in Trx f-overexpressing plants was connected to an increase in the capacity of soluble starch synthases during the light period. Taken together, these results suggest that NTRC and the ferredoxin/Trx system play distinct roles in starch turnover.
Hide AbstractGene Expression Profiling in Short‐Term Imbibition of Wheat: Tools for Dissecting of Pasting Properties of Imbibed Wheat Seeds.
Tamura, T., Akuzawa, S. & Mura, K. (2019). Journal of Food Science, 84(5), 946-953.
Germination of wheat maximizes phytochemical content and antioxidant activity while altering chemical composition, gluten content, and pasting properties. This study investigated the effect of short‐term imbibition on gene expression profiles and the physical and functional characteristics of wheat. Changes in gene expression profiles of wheat during short‐term imbibition (0, 16, and 24 hr) were evaluated by DNA microarray analysis. Gene Ontology (GO) analysis was carried out to categorize the function of genes with altered expression. Genes related to cellulose and cell wall synthesis were upregulated by imbibition for 16 hr, whereas those associated with polysaccharide catabolism and nucleosome assembly were upregulated in the subsequent 8 hr. The genes related to proteases and gluten were expressed in dry seeds but disappeared after 16 hr of imbibition. Genes encoding α‐amylase were not expressed in dry seeds whereas those encoding β‐amylase were expressed in dry seeds and downregulated by imbibition. According to quantitative real‐time PCR and enzymatic activity assay, α‐Amylase expression increased by imbibition and reached a maximum 24 hr after imbibition, with a corresponding increase in enzymatic activity. Pasting properties of flour made from wheat seeds imbibed for different times were decreased when seeds were imbibed for over 16 hr, by examination with Rapid Visco Analyzer. Gluten content did not significantly change until 24‐hr imbibition, although expression of genes encoding gliadin and glutenin disappeared by 16‐hr imbibition. The data indicated that it was possible to use 16‐hr imbibed wheat, with up to the 50% w/w replacement of nonimbibed wheat.
Hide AbstractRice malting optimization for the production of top‐fermented gluten‐free beer.
Ceccaroni, D., Marconi, O., Sileoni, V., Wray, E. & Perretti, G. (2019). Journal of the Science of Food and Agriculture, 99(6), 2726-2734.
Background: A safe method to obtain gluten‐free beer led to the use of naturally gluten‐free grains, such as rice, but the specific malting program for rice is long and requires a large amount of water, and the resulting beer showed a flat flavour profile. In this study, an optimization of the malting and brewing procedure is proposed to overcome the aforementioned issues. Different steeping conditions and kilning temperatures are considered, and a top‐fermented beverage from rice malt is obtained for the first time. Results: The malting procedure has been optimized by assessing the use of short‐time steeping as an alternate to long air rest to obtain sufficient moisture content in the green malt, saving water consumption. The malt obtained allowed a regular fermentation, as confirmed by the sensorial analysis, which did not reveal any off‐flavours. The use of a top‐fermenting yeast formed high content of higher alcohol and relatively low amount of esters. Conclusion: This study confirms the potential of rice for the production of malt and beer. The optimized malting programme allowed water saving. The production of a top‐fermented rice malt beer was a successful attempt to introduce a new flavoured product for consumption by individuals affected by coeliac disease.
Hide AbstractDi Ghionno, L., Marconi, O., Lee, E. G., Marconi, O., Rice, C. J., Sileoni, V. & Perretti, G. (2017). Journal of Agricultural and Food Chemistry, 65(23), 4777-4785.
This study was conducted to evaluate the behavior of a white teff variety called Witkop during malting by using different parameters (germination temperature and duration) and to identify the best malting program. Samples were evaluated for standard quality malt and wort attributes, pasting characteristics, β-glucan and arabinoxylan content, and sugar profile. It was concluded that malting teff at 24°C for 6 days produced acceptable malt in terms of quality attributes and sugar profile for brewing. The main attributes were 80.4% extract, 80.9% fermentability, 1.53 mPa s viscosity, 7.4 EBC-U color, 129 mg/L FAN, and 72.1 g/L of total fermentable sugars. Statistical analysis showed that pasting characteristics of teff malt were negatively correlated with some malt quality attributes, such as extract and fermentability. Witkop teff appeared to be a promising raw material for malting and brewing. However, the small grain size may lead to difficulties in handling malting process, and a bespoke brewhouse plant should be developed for the production at industrial scale.
Hide Abstract