| Content: | 22,000 Units |
| Shipping Temperature: | Ambient |
| Storage Temperature: | 2-8oC |
| Formulation: | In 3.2 M ammonium sulphate |
| Physical Form: | Suspension |
| Stability: | Minimum 1 year at 4oC. Check vial for details. |
| Enzyme Activity: | Dehydrogenase |
| EC Number: | 1.1.1.28 |
| CAS Number: | 9028-36-8 |
| Synonyms: | D-lactate dehydrogenase; (R)-lactate:NAD+ oxidoreductase |
| Source: | Leuconostoc mesenteroides |
| Molecular Weight: | 36,455 |
| Concentration: | Supplied at ~ 20,000 U/mL |
| Expression: | Recombinant from Leuconostoc mesenteroides |
| Specificity: |
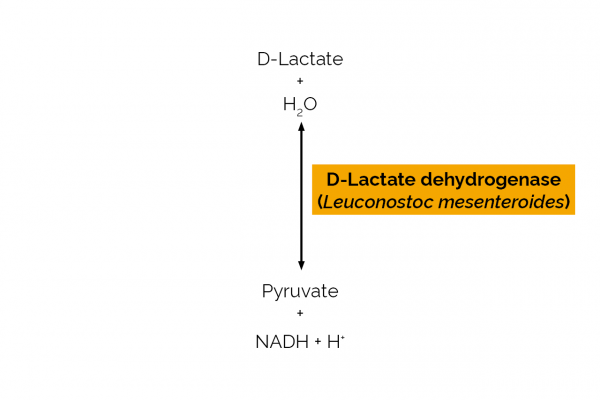
Catalyses the reaction: D-Lactate + NAD+ = pyruvate + NADH + H+ |
| Specific Activity: | ~ 1,500 U/mg (25oC, pH 7.0 on pyruvic acid) |
| Unit Definition: | One Unit of D-lactate dehydrogenase is defined as the amount of enzyme required to produce one µmole of D-lactate from pyruvic acid (0.77 mM) per minute in the presence of NADH in sodium phosphate buffer (92 mM), pH 7.0 at 25oC. |
| Temperature Optima: | 37oC |
| pH Optima: | 7 |
| Application examples: | Applications for the measurement of D-lactic acid in the food, fermentation, wine, beverage and dairy industries. |
This product has been discontinued (read more).
High purity recombinant D-Lactate dehydrogenase (Leuconostoc mesenteroides) for use in research, biochemical enzyme assays and in vitro diagnostic analysis.
Display all analytical enzymes.
Oppermann, H., Ding, Y., Sharma, J., Paetz, M. B., Meixensberger, J., Gaunitz, F. & Birkemeyer, C. (2016). Nutrition & Metabolism, 13(1), 70.
Background: Tumor cells are highly dependent on glucose even in the presence of oxygen. This concept called the Warburg effect is a hallmark of cancer and strategies are considered to therapeutically exploit the phenomenon such as ketogenic diets. The success of such strategies is dependent on a profound understanding of tumor cell metabolism. With new techniques it is now possible to thoroughly analyze the metabolic responses to the withdrawal of substrates and their substitution by others. In the present study we used gas chromatography coupled to mass spectrometry (GC-MS) to analyze how glioblastoma brain tumor cells respond metabolically when glucose is withdrawn and substituted by pyruvate. Methods: Glioblastoma brain tumor cells were cultivated in medium with high (25 mM), medium (11 mM) or low (5.5 mM) glucose concentration or with pyruvate (5 mM). After 24 h GC-MS metabolite profiling was performed. Results: The abundances of most metabolites were dependent on the supply of glucose in tendency but not in a linear manner indicating saturation at high glucose. Noteworthy, a high level of sorbitol production and release was observed at high concentrations of glucose and high release of alanine, aspartate and citrate were observed when glucose was substituted by pyruvate. Intermediates of the TCA cycle were present under all nutritional conditions and evidence was found that cells may perform gluconeogenesis from pyruvate. Conclusions: Our experiments reveal a high plasticity of glioblastoma cells to changes in nutritional supply which has to be taken into account in clinical trials in which specific diets are considered for therapy.
Hide Abstract